If you install R in a folder other than Applications, you must specify the full path. Though the R application is located in the Applications folder, the actual executable file is located within the above mentioned path. Set the R path by typing “/usr/local/bin/R” (no quotes) within the box. If installing R on a Mac: Install R in your Applications folder.Follow the links and installation directions for your operating system. Download R from the Comprehensive R Archive Network (CRAN) website. To run a Plugin that calls to the R statistical computing environment: For these plugins to work, R must first be installed and setup with the appropriate R packages. Please note that the Spade, CellOntology and FlowMeans plugins utilize the R statistical computing environment to produce results. Installing the R statistical computing environment Your plugins should now appear listed in the Plugins menu.Īs new plugins become available, they will be posted for download from the FlowJo Exchange website, which can be accessed from within FlowJo under the Plugins Menu. 
If you do not see individual plugins listed within the Plugins menu following installation, you may need to tell FlowJo the location of the plugins folder by specifying the folder’s file path in the Diagnostics section of FlowJo’s Preferences (FlowJo / Preferences / Diagnostics). (Workspace Tab –> Populations Band –> Plugins menu) Plugin actions can be accessed and initiated from within FlowJo under the Plugins Menu. Add your plugin of interest to the plugins folder on your machine, and restart FlowJo:.Follow the steps outlined by the installer and save the plugins folder to your hard drive. 
Installing Plugins in FlowJo v10.1r7+:ĭownload the latest release of FlowJo v10 for Mac or Windows These can be installed and used as shown below. 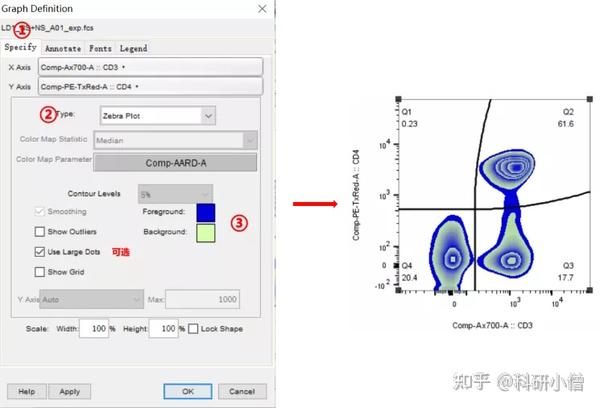
Five plugins are included with the FlowJo v10 installation package. If you run a second analysis with the same cells to get an technical error, there will be a difference from the first analysis but only because the propidium iodide has killed some of the cells! This is why beads are used to check the coefficient of variation of the actual analysis.Plugins are executable java files that extend functionality of the FlowJo application. So if you leave it on long enough it will begin to kill the cells. A classic example of this is propidium iodide, often used as a cell viability die. Running the same sample 3 times will not really tell you anything as the very presence of the probe may cause the cell to respond differently on the 2nd or 3rd analysis. Id ask your lab manager about that.Įxperimental and biological error though will have to be performed by staining independently grown cell samples and then running 3 or 4 seperate experimental repeats to get statistical error. To overcome this, most flow software has some sort of a built in coefficient of variation analysis that often involves beads. They seem to be of the opinion you dont need technical repeats in flow as you are analysing thousands (in some cases tens of thousands) of events. Your question about technical error is interesting, its an argument ive had with immunologists. Then tell it to count the percentage of cells that fall into that region. I think in the flowjo software you can set regions you want the software to count like dividing the plot into quads, polygons or whatever. There is a way of inserting stats onto your histograms or dot plots.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
